Stéphanie Bury Moné: Revealing the hidden potential of bacteria
Stéphanie Bury Moné, a lecturer at the Institute for Integrative Cell Biology (I2BC - Univ. Paris-Saclay, CEA, CNRS) and Deputy Director of Studies at the Graduate School Life, Science and Health at Université Paris-Saclay, specialises in blending different research techniques, including genomics, to reveal the hidden talents of soil bacteria, which are capable of naturally developing antibiotics. Understanding how their natural defence system works is a step forward towards discovering and understanding antibiotics for human medicine.
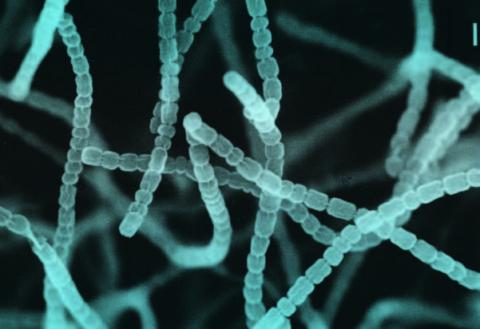
Born from the merger of several laboratories on the Université Paris-Saclay campus, I2BC is one of the largest research units in biology at the university. Its speciality is studying the cell and its interactions. Within the genome department, Stéphanie Bury Moné is part of the Conformation and segregation of the bacterial chromosome team, led by Frédéric Boccard. More specifically, she studies the genome of bacteria of the Streptomyces genus. Relatively unknown to the wider public, these bacteria are widespread in soil and can make a significant contribution to human medicine. “A third of antibiotics used clinically come from these bacteria. Our goal is to reveal the natural biocomponent potential of micro-organisms that produce them to defend themselves and continue to thrive.”
In search of the hidden metabolism
“Studying the genome of Streptomyces reveals the presence of many more potentially coded molecules than we thought. However, we still can’t find the conditions in which the bacterium will produce them,” explains Stéphanie Bury Moné, who nowadays is focused on genomic dynamics. “By trying to understand how gene expression is regulated and how these large clusters code the biosynthesis of antibiotics will be activated or suppressed, we will be able to control them and discover new ones.”
Dynamic observation of genomes, which involves ‘capturing’ and comparing contacts within chromosomes, is a team speciality that is recognised internationally. It carries out this work using the I2BC sequencing platform. “Analysis of the ‘core’ genome can be used to identify the behaviour of ultraconserved elements compared to very variable parts.” This method combines bioinformatics, to compare species between one another, and analyse the evolution of the genome over time, and genomics, to precisely clarify the conditions of expression of the genome during its development cycle in relation to the three-dimensional architecture of DNA. This different approach aims to define how bacteria remodel their chromosome upon antibiotic production.
Clusters of antibiotics
In 2021, this lecturer and her team were able to reveal the dynamics of remarkable genome of the Streptomyces bacteria. “Although the majority of bacteria have a circular genome, that of Streptomyces is linear. Its centre is well conserved and at each end, we can see several genomic clusters, including those that code specialised metabolisms.” When the bacterium enters the stationary phase (starvation), it compresses its genome and forms a new structure where expression of all genetic information that is needed to produce an antibiotic will happen at the highest level. This initial description of the dynamics of antibiotic production was the subject of an article in Nature Communications.
Although Stéphanie Bury Moné admits to having thought about the potential practical applications of this research, this, as of yet, is still fundamental experimental biology. “I’m waiting to reach a level of sufficient progress in the elucidation of the rules that govern the genomic organisation of Streptomyces bacterium. We need to be able to better control the stability conditions of strains over time in order to optimise antibiotic production and exploit genomes for industrial purposes.” To do this, within the framework of the STREPTOMICS project that she is leading, she is actively involved in the development of new genomics techniques capable of targeting DNA-associated proteins that regulate the dynamics of bacterial development. “One day, we might be able to mutate them, overexpress them and control them.”
A prestigious career
Stéphanie Bury Moné’s passion for micro-organisms goes back to her childhood. As a student, she enrolled at ‘Université de Nice’ after a year of a ‘classe préparatoire’ (2-year preparatory course for a competitive exam) in advanced maths and biology to do a first year of biology and biochemistry. She passed the competitive entrance exam for the ENS Cachan (today, ENS Paris-Saclay), one of the few institutions that provided a preparatory course for the biochemistry & biological engineering ‘agrégation’ competitive exam. She succeeded the following year. This success opened the door to the Institut Pasteur, where her dream came true: doing a second year of a Master’s degree and then a PhD in microbiology, supervised by Hilde de Reuse. The Belgian researcher, a specialist in Helicobacter pylori bacteria, became her mentor and definitively gave her a taste for research. In 2003, Stéphanie Bury Moné defended her thesis about the resistance of pathogenic bacterium to gastric acidity. She is one of the first people to comprehensively analyse the expression of genes of these bacteria in response to acidity.
The genome, from one cell to another
With her desire to explore the microbial world, Stéphanie Bury Moné turned to viruses. She joined Jean-François Mouscadet’s team at the Laboratory of Biology and Applied Pharmacology (LBPA – Univ.Paris-Saclay, ENS Paris-Saclay, CNRS) to study HIV. She divided her time between research and teaching as an ‘Attaché temporaire d’enseignement et de recherche’, ATER, (graduate teaching assistant). She was awarded the role of lecturer at the Institute of Genetics and Microbiology. With Philippe Bouloc, she worked on the Escherichia coli bacteria’s response to stress for a while, before returning to work on HIV at LBPA. In 2012, she obtained her ‘habilitation à diriger des recherches’, HDR, (authorisation to supervise the research of others) in virology. After nine years spent on studying HIV, Stéphanie Bury Moné went back to studying bacteria in collaboration with Sylvie Rimsky, a researcher at the ‘Collège de France’. “No matter the cellular model, the central theme of my research remains the study of genomes and their expression. The different knowledge I have acquired is mutually enriching, and is an asset.”
Combining research and learning
Stéphanie Bury Moné has been the co-director of the Master 2 in Fundamental Biology since 2015, and was promoted to Professor at the Faculty of Science at Université Paris-Saclay in 2016. This event was the starting point for her commitment to developing the Graduate School for Life Sciences and Health at Université Paris-Saclay, of which she became a Deputy Director of Studies in 2020. “This Graduate School was created on the basis of a merger of various models of Université Paris-Saclay institutions, whose biology-health network already existed. We brought together the fields of biology, bioinformatics and medicine to build a programme of teaching that ranges from the molecule to synthetic biology, including biochemistry, oncology, immunology, biotherapy, microbiology, neuroscience, human physiology, development, bioinformatics, genetics and genomics.” The Graduate School has 1,500 students (1,000 Master’s students and 500 PhD candidates) and 2,000 permanent staff.
Stéphanie Bury Moné devotes as much time to research as she does to teaching. “I’m lucky to have the most wonderful job.” She is particularly committed to her students, and believes that it is her mission to lead them “to the very end of their project, by highlighting their skills. They should be proud of what they have learnt.”
